Pages about Jmol-specific topics (This link runs a search for all pages within Proteopedia that have titles beginning with "Jmol".It is usually convenient to gather commands in script files. You can run a script file by dragging it and dropping it into the molecular graphics window of Jmol. You are now ready to enter Jmol commands into the Jmol Script Console. The names of command script files should always end. Command script files must be edited with a plain text editor. You are now ready to enter Jmol commands into the Jmol Script Console. Open Jmol's File menu (above the black window), and Open or Open Recent.Enter the command "load ?", and a file picking dialog will appear.Drag the file and drop it into the black Jmol window.The "=" tells Jmol to get the file from the Protein Data Bank. Enter the command "load =1d66" in the white window (without typing the quotation marks).If you do not have a downloaded copy of the PDB file:.There are several ways to load a PDB file:. 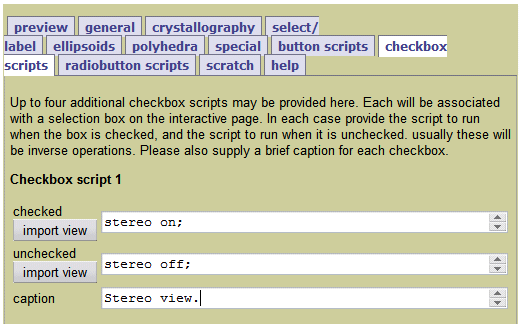
In java windows, the usual Apple Cmd-C and Cmd-V don't work! You can also select blocks of text with your mouse, then drag and drop them. This no longer seems necessary for OS X: Copying and pasting: To copy or paste commands in the white Jmol Script Console window, use Ctrl-C and Ctrl-V, even if working on an Apple Mac OS X!.Now you are ready to enter commands (see the Jmol Interactive Scripting Manual). Open the Jmol Script Console: If the white "Jmol Script Console" window does not appear, use Jmol's File menu (at the top), Console to open it.Run the Jmol application: Double-click Jmol.jar in your working folder, and a black window will appear titled "Jmol".Alternatively, or if you still get an Out Of Memory error, you can give Jmol more memory to work with.You can check Jmol's java memory usage by clicking on Jmol (in either the applet or the application), then on the main menu, About, System. Close all browser windows or tabs that contain the Jmol applet. If you skip this step and Jmol gives you an "Out Of Memory" error, then come back and do this step. This is not necessary if your molecule(s) are modest in size (up to ~30,000 atoms). This is probably no longer important: Free java memory if your project involves large molecules or assemblies.Double click the zip file to unzip it, and copy the file Jmol.jar (one of a large number of files in the zipped download). Go to Jmol.Org, select Download (upper right), go to the Jmol Downloads Page, and download the current version as a binary.zip file. Put Jmol.jar in your working folder: Create a folder (directory) on your computer in which to work.You can download and install it easily at. The Jmol application requires Java (free) in order to run.For some advanced projects, you may wish to use the Jmol standalone application (no web browser involved once it is downloaded). Pages in Proteopedia use a javascript form of Jmol called JSmol, which operates automatically within a web browser. 
Jump to: navigation, search proteopedia link proteopedia link 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
